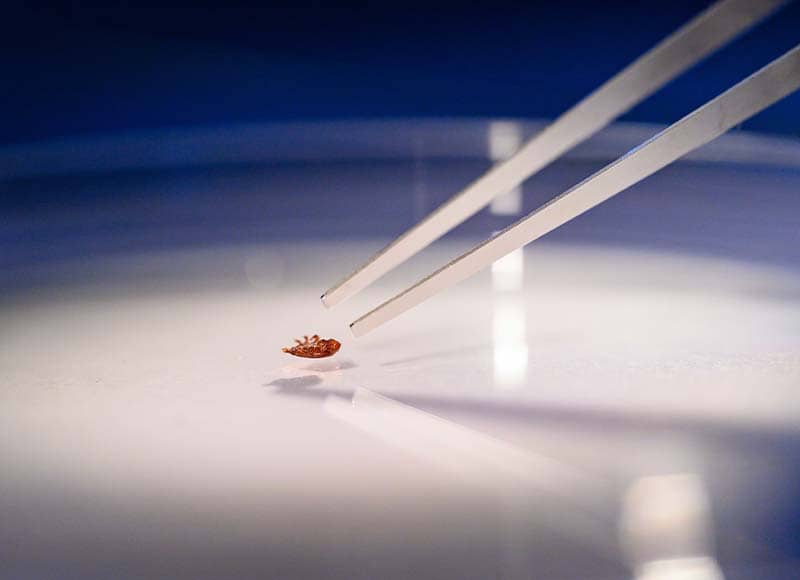
Benedicte Fustec uses tweezers to position a bloated tick on its microscope stage for an elaborate photo shoot. Its blood-red color gives away the tick’s favorite meal.
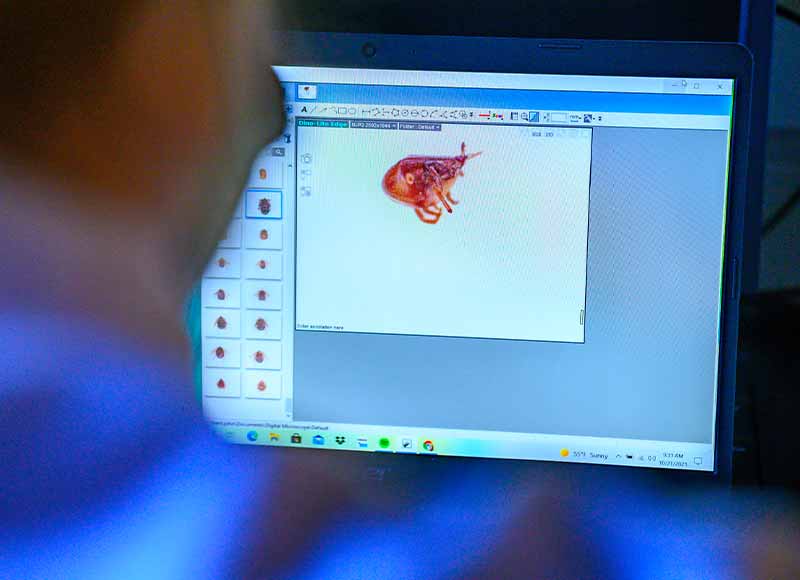
A postdoctoral researcher in the biology lab of Nicole L. Achee and John P. Grieco, Fustec is taking Vogue-quality pictures of known ticks in order to train an experimental artificial intelligence program how to identify the correct tick species. Only a few specialists in the world can make a reliable identification by sight, but the AI program will learn the morphological features (such as mouth size or leg color) that distinguish different tick species by processing up to 100 pictures during each of its three life stages for both male and female specimens.

That knowledge — especially if it’s known quickly and widely — can inform scientists about the link between species of tick and presence of tick-borne pathogens (viruses, bacteria, parasites) with the potential to become the next global pandemic, and also allow for better risk projection and early prevention, which are the key to any containment plan.
It’s a long road from bug pictures to a worldwide early detection system that can identify the next COVID-19. Achee and Grieco recently started a 15-month first phase of a project called REDI-NET, which stands for Remote Emerging Disease Intelligence Network.
“We want to improve the accuracy and timeliness of the ‘data-to-decision’ pipeline,” Achee said.
Funded by the U.S. Army Medical Research and Development Command as a way to protect American troops and their dependents wherever they are stationed, as well as communities in these locales, the goal is to provide actionable data to decision-makers in military, government or health sectors. A process that was unreliable in a field lab or that could take weeks for samples to be sent overseas could be available online almost immediately through a new data portal custom-built and housed at Notre Dame’s Center for Research Computing.

“We want to improve the accuracy and timeliness of the ‘data-to-decision’ pipeline.” —Nicole L. Achee, Lead Principal Investigator
“The big picture is that we’re going to get more real-time data,” said Grieco. “You can develop risk assessments for when a tick population becomes large enough to spill a pathogen into humans. These types of things are going to, hopefully, allow us to make better predictions at a faster rate, and get on top of these type of epidemic situations before they get out of control.”
The next pandemic is not likely to start blocks from the Centers for Disease Control in Atlanta. The unknown pathogen will instead make the jump to humans in a remote location where people live in close contact with animals.
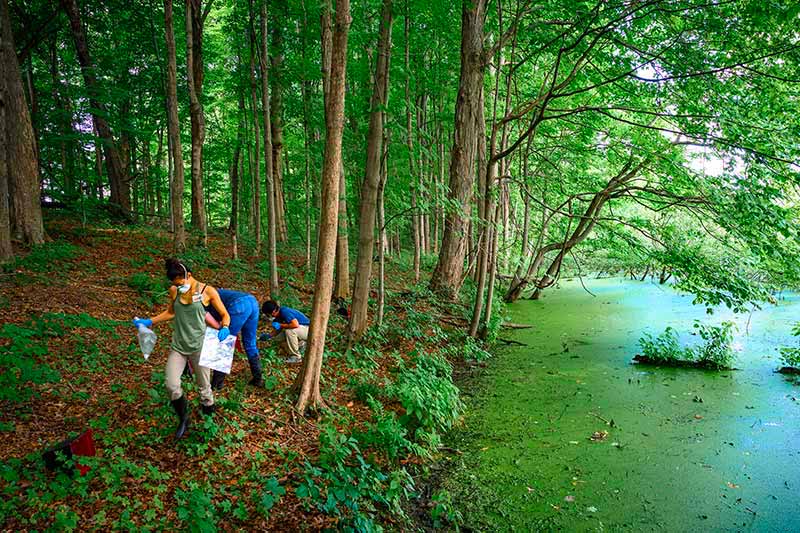
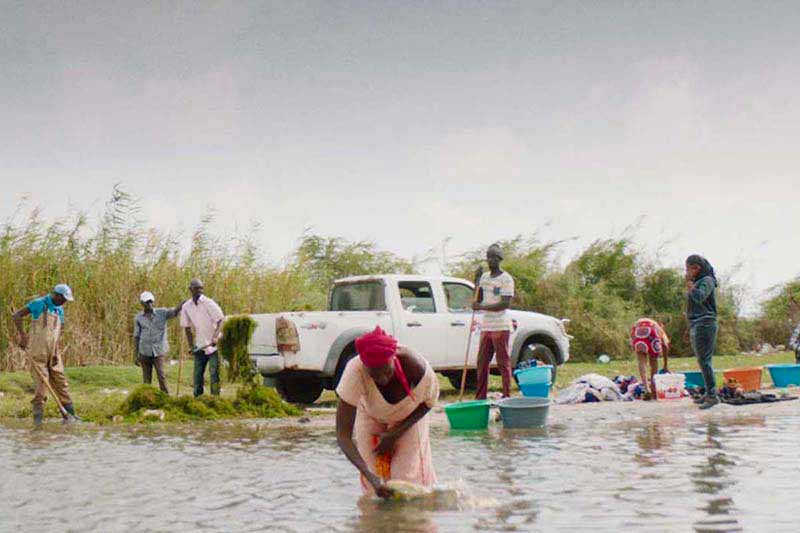
Grieco and Achee have worked for years with labs across Latin America, Africa and Asia. The first phase of REDI-NET begins with creating standard procedures for collecting reliable data and formatting it into a computer platform capable of going beyond the basic information to features like complex data filters and data visualization. Later phases will expand partnerships across geographies and ecologies to include military, health and government labs around the world that could input their local field research.
“One of the challenges in disease surveillance is the capacity for testing samples in the locations where these circulating pathogens exist,” Achee said. “With this system, we can integrate the pathogen data with ecology data from where we sample to better inform partners who are responsible for conducting control activities or mitigating the risk of pathogens infecting humans or animals in that environment.”
In early November, the REDI-NET team presented an update on their progress to a group from the Military Infectious Diseases Research Program ( MIDRP), which manages the defense funding for the project. The director, Col. Stuart Tyner, is a former offensive lineman from the Notre Dame football team under Coach Lou Holtz.
“We know that biological vectors (organisms that transfer pathogens) are key to limiting or preventing disease for our warfighters,” Tyner told Achee and Grieco. “I think that there are some opportunities with vector-based research to solve our problems.”
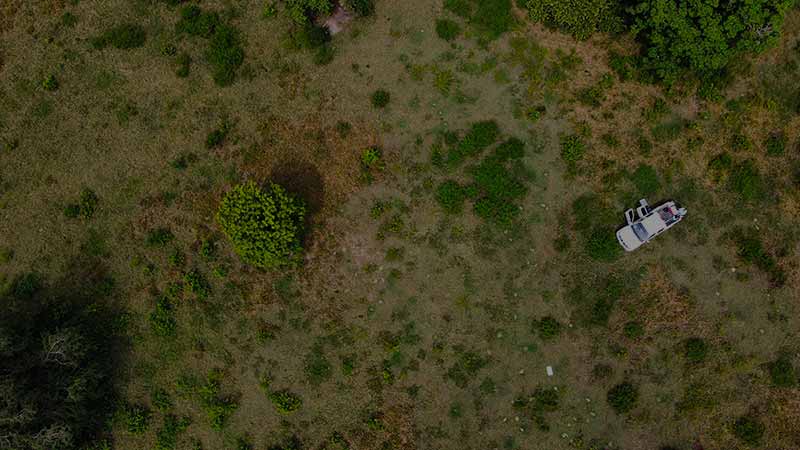

In REDI-NET’s lab in the Galvin Life Science Center, Donovan Leiva holds up a specialized piece of cloth and explains to the MIDRP group how to drag a field for ticks.

“Through this work, I’ve seen opportunities to improve health care and research in my country, and I hope to contribute.” —Donovan Leiva, Graduate Student
Keough School of Global Affairs
Leiva, currently a graduate student in the Keough School of Global Affairs, is from Belize and began volunteering in Achee and Grieco’s Belize Vector and Ecology Center (BVEC) just before graduating from the University of Belize with a biology degree. He advanced from intern to research assistant to acting manager over four years at BVEC, involved in everything from tick collection and hosting community health education events to training vector-control government officers, supervising Notre Dame students and speaking at international scientific conferences.
Last year, Leiva decided to further his education with a master’s degree at Notre Dame, and Grieco and Achee hired him to work part-time alongside three undergraduate researchers on this end of their operation. “Through this work, I’ve seen opportunities to improve health care and research in my country, and I hope to contribute,” Leiva said.
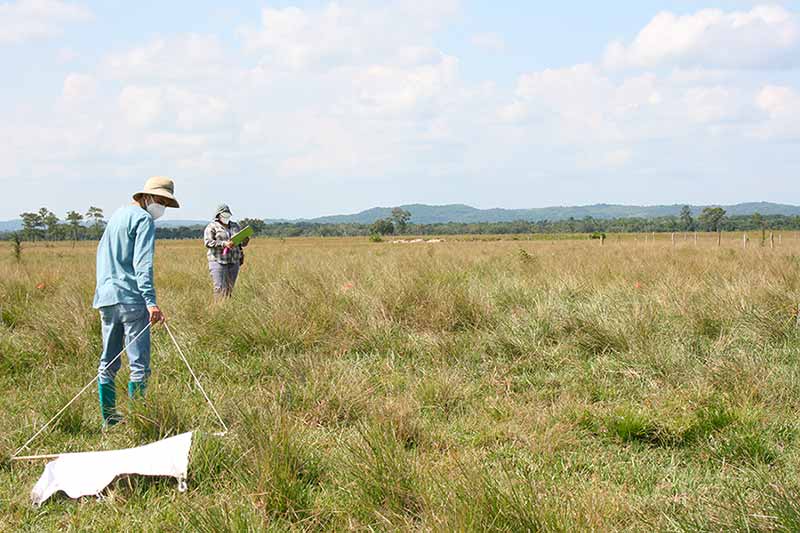

That kind of local participation and connection-forming help Achee and Grieco build partnerships in their fieldwork around the world. The married couple have been working together to better understand disease vectors for more than 25 years. They have joint appointments in the Department of Biological Sciences and the Eck Institute for Global Health.
Their experience helps them understand the challenging conditions under which remote labs operate. Phase 1 of the REDI-NET program aims to establish standard operating procedures, training materials and new technology that will allow vector and disease data to be compared across locations and time.
For instance, the program will use tablets and a custom-designed mobile app to input data on site in the field, whether there is an internet connection there or not. Information fields about collection location and environmental conditions include drop-down menus to ensure data gets formatted in the same manner everywhere.
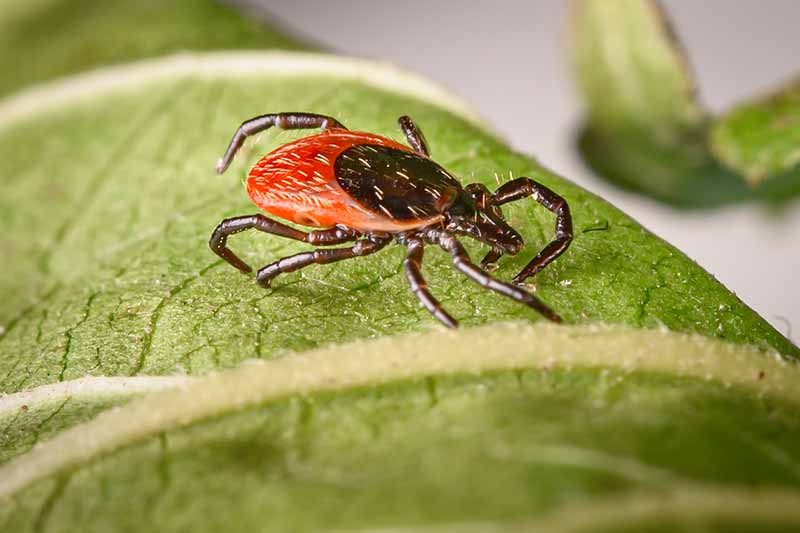

“Different tick species carry different pathogens and have different ecologies,” Achee said. “That means they might feed on different types of animals that might live in different types of environments, maybe a savannah versus a mountainous area versus a swamp.”
Besides ticks, the program will collect and process leech, water and sediment samples. Mosquitoes, the best-known vector, may come later because most other programs focus solely on mosquitoes. The goal of the first phase of REDI-NET is to establish a foundation for collecting samples in a standardized way and strengthen capacity for testing in field labs.
“It’s not that pathogen surveillance is something new,” Achee said. “What we’re trying to do is enhance a system that has gaps.”
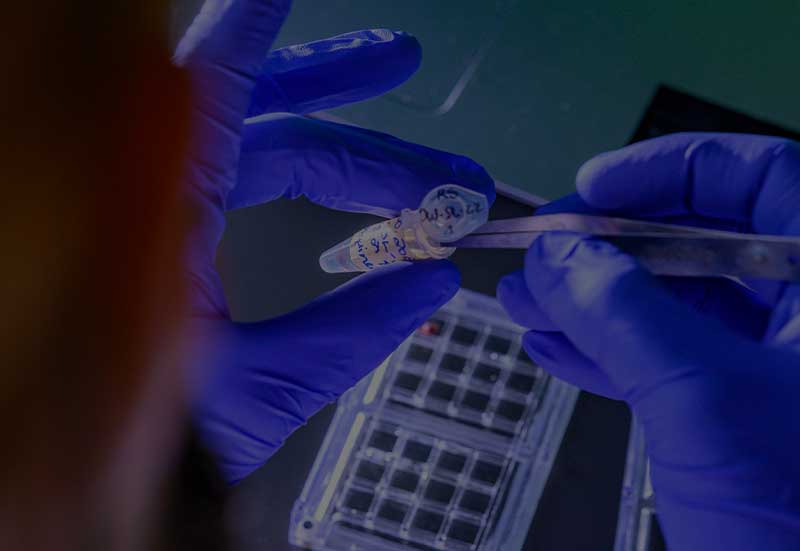
The machine that Fustec is training to identify tick species was developed by Vectech, a biotech company that has already shown success with AI learning for mosquito identification. It can take a picture of 12 ticks at a time, so it will speed up a process that can be slow in the field.
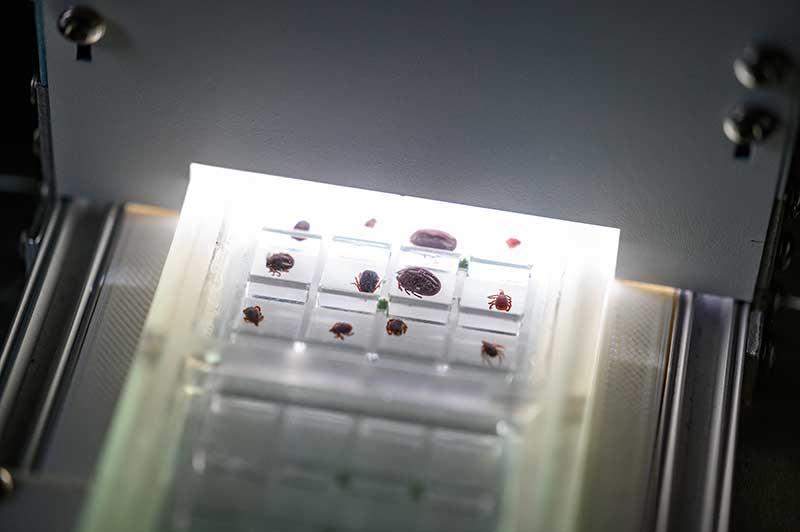

Labs at Notre Dame, the Naval Medical Research Center and the Walter Reed Biosystematics Unit are considered Gold labs within the REDI-NET system because of their expertise and responsibility to validate outcomes. Field labs in Florida (wetlands), Belize (tropical forest) and Kenya (tropical grasslands) are identified as Silver labs. All will have the ability to test for pathogens.
The second phase of REDI-NET will expand the field sample portfolio, improve testing functionality, validate surveillance models and train partners in the Defense Department.
Each tick, leech, water and soil sample will be given a unique barcode that will link field and lab data about that sample and quickly identify the storage location information loaded into the data portal. Then Fustec demonstrated how a tick will be ground up so that another machine can separate out its DNA and detect the presence and type of pathogen it carries — anything from dengue fever or coronavirus to novel or little-known pathogens that may cause human diseases.


“I could probably list off 20 diseases circulating right now — that you’ve never heard of, that are right on our doorstep — that are mosquito or tick-borne,” Grieco said. “Mayaro fever (an arbovirus related to dengue fever) is spreading from South America into Mexico and potentially onto our southern border as we speak. We don’t know where the next threat is going to come from. This program allows us to identify potential locations.”
Ebola. Swine flu. West Nile virus. Zika. Lyme disease. Avian flu. SARS. These diseases are known as zoonotics because they circulate regularly in animal populations like bats or monkeys but can jump over to humans and cause deadly outbreaks. What other people fear as the future zombie apocalypse, Grieco and Achee see as an opportunity.

“I could probably list off 20 diseases circulating right now — that you’ve never heard of, that are right on our doorstep — that are mosquito or tick-borne.” —John P. Grieco, Principal Investigator
“There is the potential for having the work done on these pathogens here at Notre Dame,” Grieco said. “A lot of these pathogens, we don’t even know they exist at this point in time, but other researchers can come here. And we have people across campus that can begin looking at maybe vaccine development, therapeutic development.”
The expanding human population has been encroaching more and more into habitats where they are coming into contact with animals that have been living naturally with these pathogens.
“There are a number of different routes pathogen infection can take,” Achee said. “When people are interacting with animals, or vectors that previously fed on these animals, they can become exposed and initiate spillover into the human population. That’s when mutations can take hold, allowing the pathogen to become more adapted to a human host, and more easily spread.”
Achee could be describing the origin of the coronavirus pandemic, where scientific evidence is building that the virus jumped from a live-animal market to a human exposed in Wuhan, China, and then spread rapidly to other people through airborne respiration.

Brad Sandberg, a research programmer in the Center for Research Computing, walked the MIDRP group through the data portal his team is building for REDI-NET. He displayed examples of sampling site maps that identify each specimen, training videos and data collection sheets displayed on the tablet. He also showed a pathogen heat map that can help leaders visualize danger hotspots or forecast developing threats — based on mathematical models built by Alex Perkins, an associate professor in the Department of Biological Sciences.
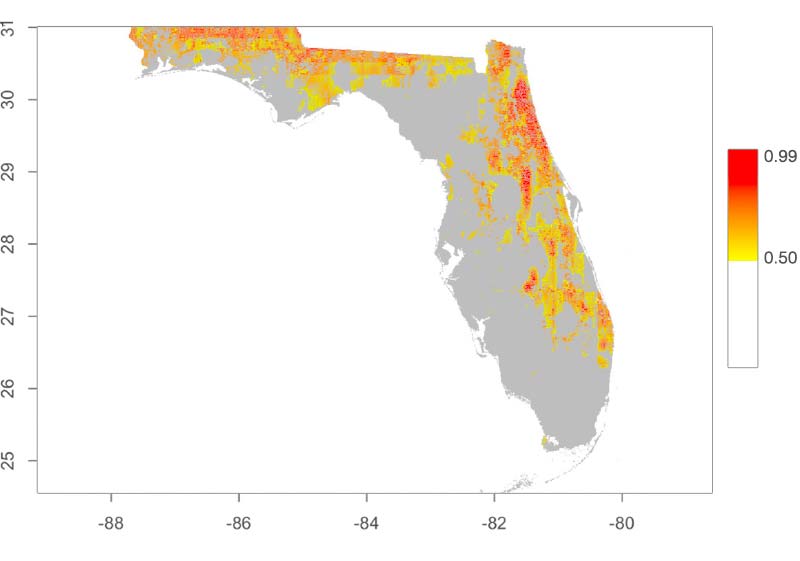
When Col. Tyner asked about aggregating data for a worldwide analysis, Achee said those questions are essential because the needs of the end-users can drive the custom creation of the data portal. “That’s why it’s so important that we developed a REDI-NET standardized approach to data collection and specific data management platform and didn’t try to adapt another platform,” she said. “We envision the user filtering for whatever they want — say, tick and pathogen spread in Cambodia.”
Phase three of the project aims to incorporate the REDI-NET system into armed forces and university labs collecting pathogen or disease vector information across the world. The sample type and data set will eventually grow in a standardized and exponential way. Authorized users will be able to log into the data portal from anywhere and retrieve whatever information they need from the cloud-based portal.
“It’s really about scaling up,” Achee said. “What we want to do is represent a global center of excellence. We want to serve as a conduit for training on field sampling, laboratory testing and data standardization, for data warehousing and modeling for stronger outputs in disease surveillance.”
Achee and Grieco don’t claim they can stop the next pandemic, which most experts expect to occur with increasing regularity in the future. But REDI-NET is being implemented to provide early warning.
“We will be better prepared the next time something happens,” Grieco said. “Maybe we can curtail the overall global impact. Stopping it? These are hard to stop, but definitely being much more prepared for the next outcome.”
This work is supported by the U.S. Army Medical Research and Development Command under Contract No.W81XWH-21-C-0001. The views, opinions and/or findings contained in this report are those of the author(s) and should not be construed as an official Department of the Army position, policy or decision unless so designated by other documentation.


